Insets and panels¶
Panel axes¶
It is often useful to have narrow “panels” along the edge of a larger
subplot for plotting secondary 1-dimensional datasets or summary statistics.
In ProPlot, you can create panels by passing a location (e.g., loc='r' or
loc='right') to the panel or panel_axes
methods. The resulting axes are instances of CartesianAxes.
To generate “stacked” panels, simply call panel more
than once. To include panels when centering spanning axis labels and super
titles, pass includepanels=True to Figure. Panels
do not interfere with the tight layout algorithm and
do not affect the subplot aspect ratios.
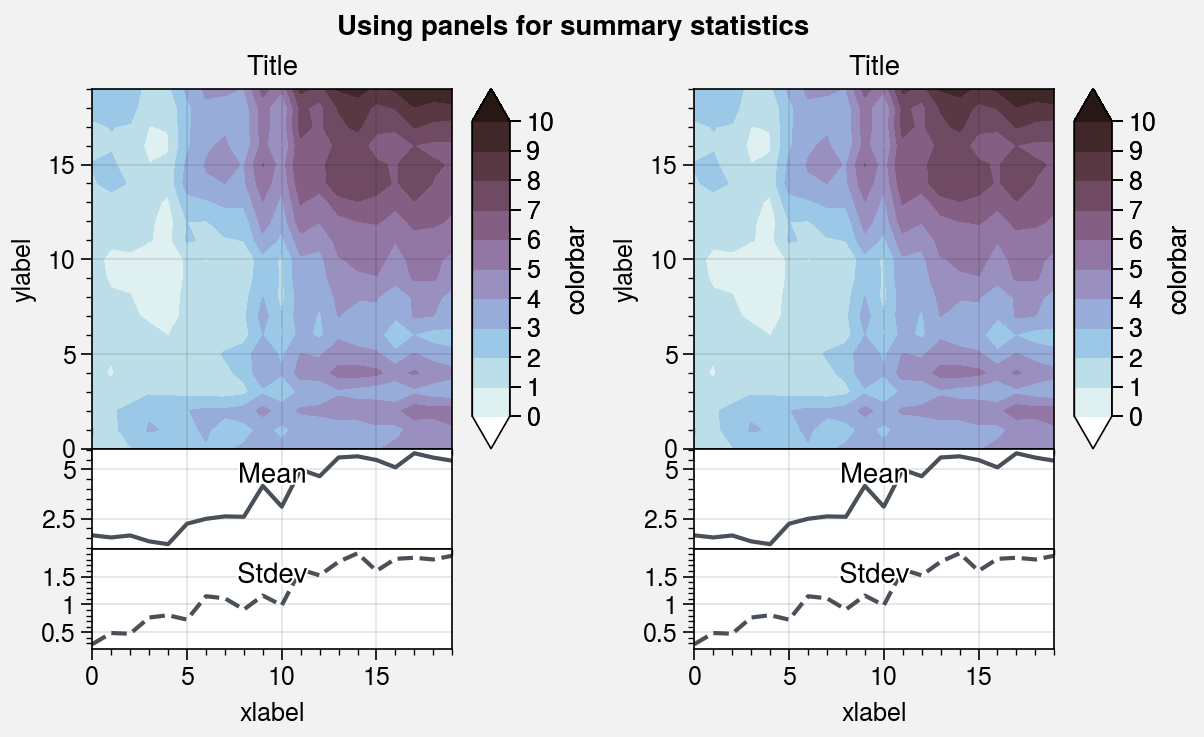
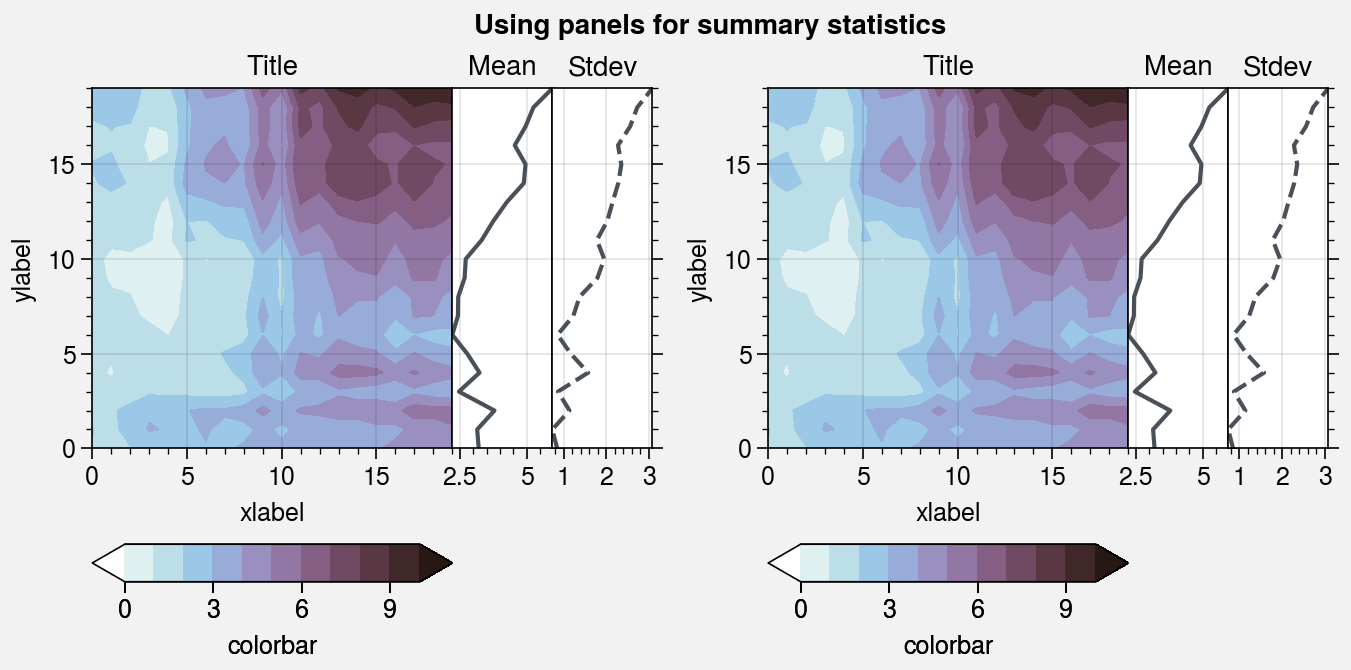
In the first example below, the panel distance from the main subplot is
manually set to space=0. In the second example, the distance is automatically
adjusted by the tight layout algorithm.
[1]:
import proplot as pplt
import numpy as np
state = np.random.RandomState(51423)
data = (state.rand(20, 20) - 0.48).cumsum(axis=1).cumsum(axis=0)
data = 10 * (data - data.min()) / (data.max() - data.min())
# Stacked panels with outer colorbars
for cbarloc, ploc in ('rb', 'br'):
fig, axs = pplt.subplots(
refwidth=1.8, nrows=1, ncols=2,
share=0, panelpad=0.1, includepanels=True
)
axs.format(
xlabel='xlabel', ylabel='ylabel', title='Title',
suptitle='Using panels for summary statistics',
)
# Plot 2D dataset
axs.contourf(
data, cmap='glacial', extend='both',
colorbar=cbarloc, colorbar_kw={'label': 'colorbar'},
)
# Get summary statistics and settings
axis = int(ploc == 'r') # dimension along which stats are taken
x1 = x2 = np.arange(20)
y1 = data.mean(axis=axis)
y2 = data.std(axis=axis)
titleloc = 'upper center'
if ploc == 'r':
titleloc = 'center'
x1, x2, y1, y2 = y1, y2, x1, x2
# Panels for plotting the mean. We make two panels at once and plot data
# on both panels at once by calling functions from SubplotsContainers.
# More realistically, you would plot data on each panel one-by-one.
space = 0
width = '4em'
kwargs = {'titleloc': titleloc, 'xreverse': False, 'yreverse': False}
paxs = axs.panel(ploc, space=space, width=width)
paxs.plot(x1, y1, color='gray7')
paxs.format(title='Mean', **kwargs)
# Panels for plotting the standard deviation
paxs = axs.panel(ploc, space=space, width=width)
paxs.plot(x2, y2, color='gray7', ls='--')
paxs.format(title='Stdev', **kwargs)
[2]:
import proplot as pplt
fig, axs = pplt.subplots(refwidth=1.5, nrows=2, ncols=2, share=0)
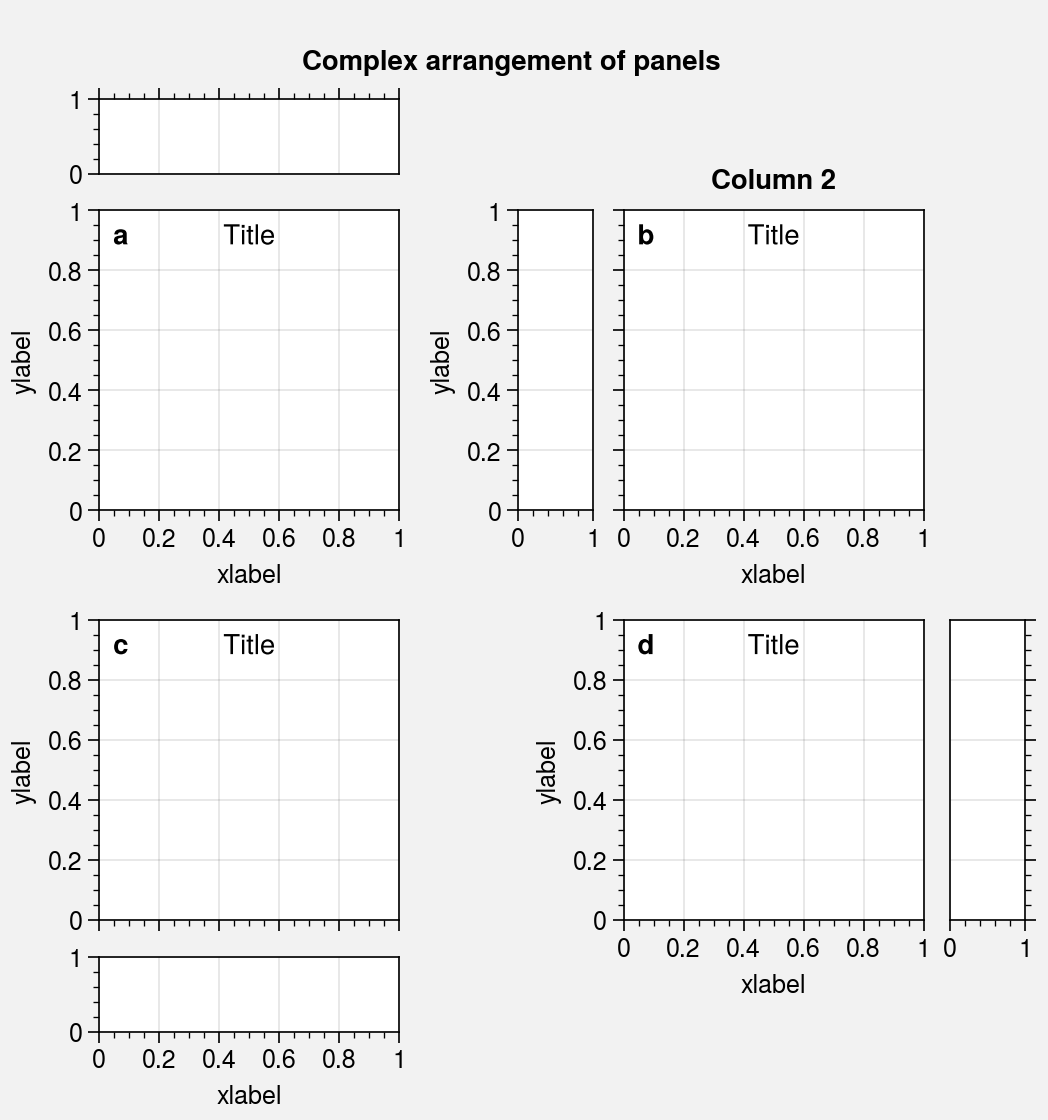
# Demonstrate that complex arrangements of panels does
# not mess up subplot aspect ratios or tight layout spacing
axs.format(
xlim=(0, 1), ylim=(0, 1),
xlabel='xlabel', ylabel='ylabel',
xticks=0.2, yticks=0.2,
title='Title', suptitle='Complex arrangement of panels',
toplabels=('Column 1', 'Column 2'),
abc=True, abcloc='ul', titleloc='uc', titleabove=False,
)
for ax, side in zip(axs, 'tlbr'):
ax.panel(side, width='3em')
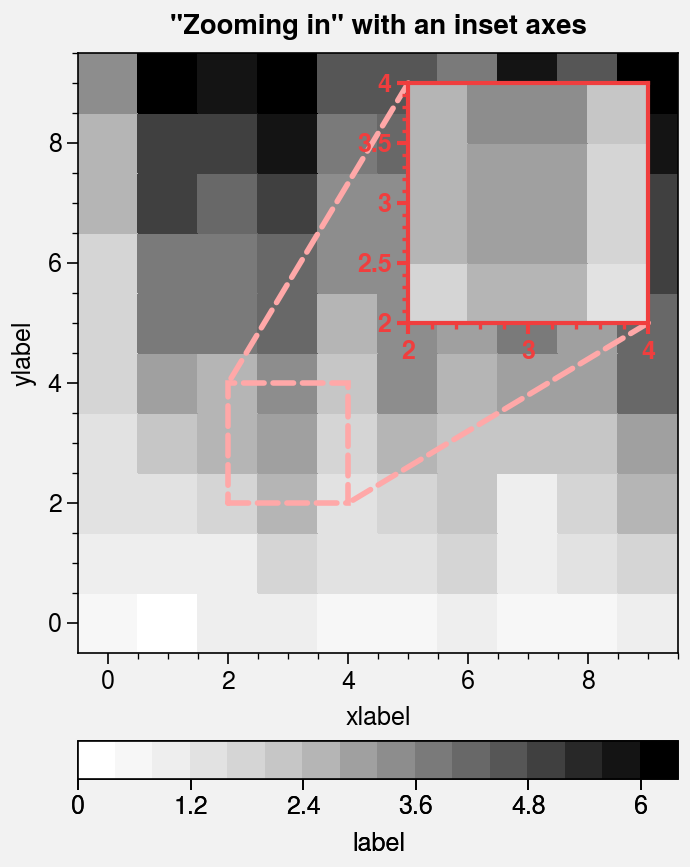
Inset axes¶
Inset axes
can be generated with the inset or
inset_axes command. By default, inset axes
have the same projection as the parent axes, but you can also request
a different projection (e.g., ax.inset(bounds, proj='polar')).
Passing zoom=True to inset draws “zoom indication”
lines with indicate_inset_zoom when the axes are both
Cartesian, and ProPlot automatically updates the lines when the axis limits of
the parent axes change. To modify the line properties, simply use the zoom_kw
argument.
[3]:
import proplot as pplt
import numpy as np
# Sample data
N = 20
state = np.random.RandomState(51423)
x, y = np.arange(10), np.arange(10)
data = state.rand(10, 10).cumsum(axis=0)
# Plot data
fig, ax = pplt.subplots(refwidth=3)
m = ax.pcolormesh(data, cmap='Grays', levels=N)
ax.colorbar(m, loc='b', label='label')
ax.format(
xlabel='xlabel', ylabel='ylabel',
suptitle='"Zooming in" with an inset axes'
)
# Create inset axes representing a "zoom-in"
iax = ax.inset(
[5, 5, 4, 4], transform='data', zoom=True,
zoom_kw={'color': 'red3', 'lw': 2, 'ls': '--'}
)
iax.format(
xlim=(2, 4), ylim=(2, 4), color='red7',
linewidth=1.5, ticklabelweight='bold'
)
iax.pcolormesh(data, cmap='Grays', levels=N, inbounds=False)
[3]:
<matplotlib.collections.QuadMesh at 0x7f63fc576a30>